

BackgroundCnidarians are often misidentified because they can appear differently in various environments. In order to learn more about the evolution of different cnidarian species, the Cytochrome Oxidase subunit 1 gene was chosen for investigation. This gene codes for a membrane protein involved in the electron transport chain, which helps to convert stored energy into ATP energy, which is usable by cells. In the below diagram, cytochrome oxidase is the third membrane complex. Cytochrome oxidase is essential to aerobic respiration, making it universal to cnidarians. |

|
|
The National Center for Biotechnology Information's primer-BLAST was used to design PCR primers that would copy portions of the COX1 gene. BLAST stands for Basic Local Alignment Search Tool; it is like a search engine for protein and nucleotide sequences stored in the NCBI databases. The "nr" or non-redundant database was chosen, because it only offers results in which the organisms have identical sequences. A primer-BLAST was run for Sarcophyton auritum, with these results limited to sequences also shared with Acropora, Ricordea florida, and Anemonia majano. This assured that the primers would be universal for a wide variety of cnidarians. The sections of DNA that continually showed up unchanged in the Cytochrome oxidase gene were made into forward and reverse primers. Primers copying segments of DNA of about 200 base pairs were chosen. |
|
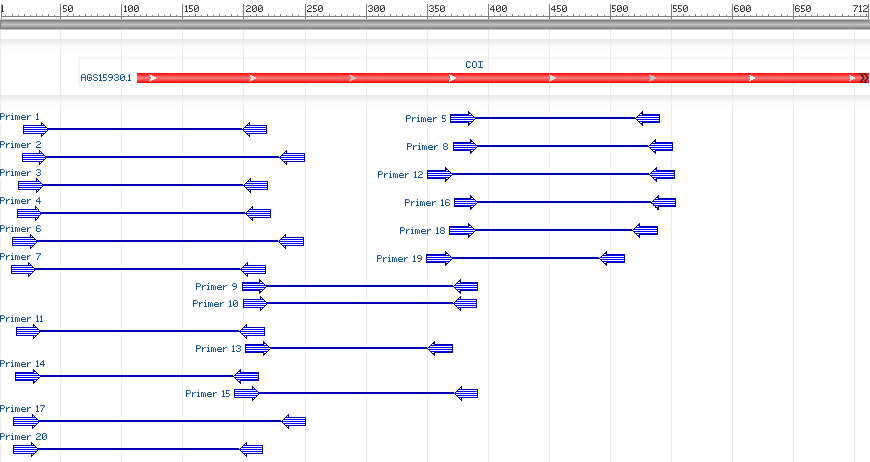
From this BLAST, three sets of primers were chosen. The first set had a forward primer of (5'-3')TTACTTTGCTGACGCACCCA and a reverse primer of (5'-3')CTCAAAGCTGTCCCTGCCAT with a length of 204 base pairs. The second had a forward primer of (5’-3’) AATGGCAGGGACAGCTTTGA and a reverse primer of (5'-3')GTCGGGCGCACCAATCATAA with a length of 193 base pairs. The last set had a forward primer of (5'-3')TATGATTGGTGCGCCCGAC and a reverse primer of CATATCCACTGCTCCCCCTG with a length of 181 base pairs. The image below shows the results of the primer-BLAST. Primer sets 3, 9, and 16 were chosen for research. |